All of our software tools are the result of close, effectiveĬollaborations with the genetic community. FAQ about confidence scores (confidence score)īioSoft provides effective, biologist friendly, easy-to-use sequence assembly and analysis software tools to meet the ever changing needs of today’s genetic/medical researcher and diagnostician.We strongly recommend you to use the SCF format instead of ABI. This is very important for companies storing large amounts of chromatogram files. In addition, the SCF format was designed to be easily compressed (using programs Not only that SCF format do not have the above-mentioned bug, but it offers many other additional features. Why the SCF format is better than ABI format? When DNA Sequence Assembler encounters a corrupted chromatogram, it will automatically try to restore the data and it will write a report (in the sameįolder where the file is located) for that corrupted chromatogram file. It can be considered an obsolete format because it has problems in storing large sequences/chromatograms. Some of my ABI chromatogram files are corruptedĪBI format was invented long time ago. low confidence in bases assignment (low confidence score) the overall amplitude of the signal is small the difference between noise and signal is very small some homopolymeric stretches are not well resolved However, if you assemble a good chromatogram and a poor one, DNA Baser will know how to extract the useful information from he good chromatogram and give less trust to the bad chromatogram. Usually, poor chromatograms enter unwanted ambiguities in your contig. A low bar means that the base call cannot be trusted. A high bar means that the base call can be trusted. These bars represents the confidence scores. 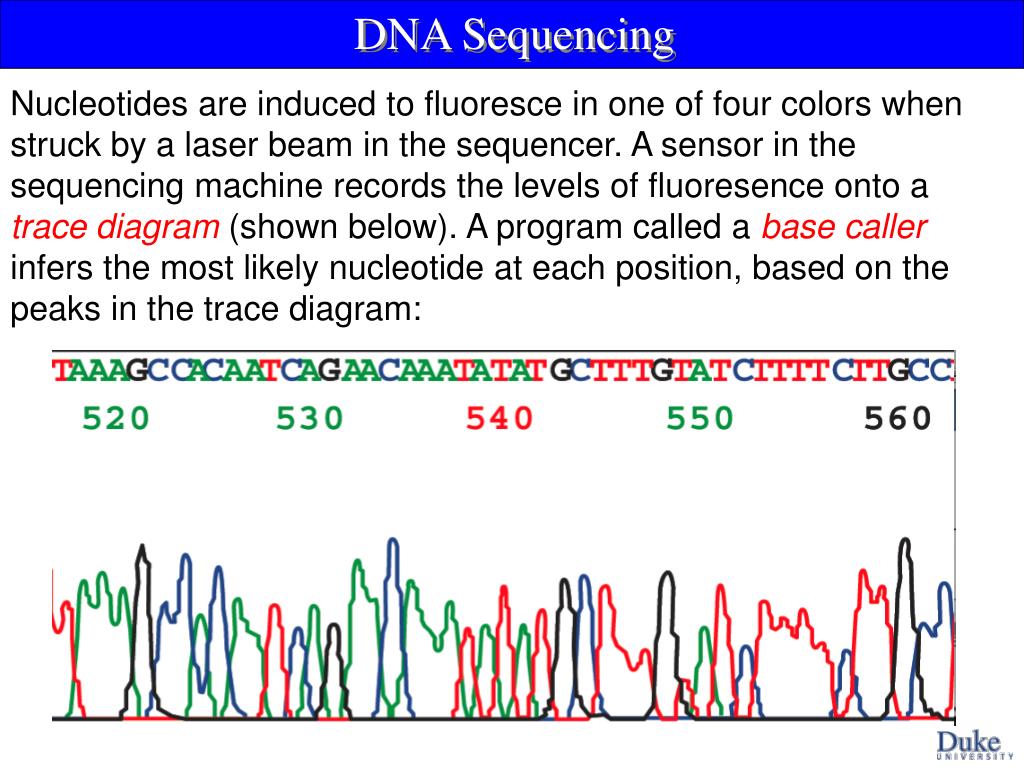
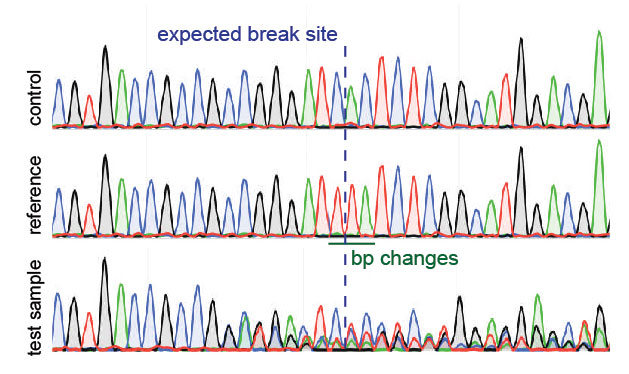
Just look at the green bars displayed above each base. With DNA Baser it is easy to make the difference between a good (trusted) chromatogram and a poor one. Please see the Download page for details. To open a chromatogramįile just double click it in Project Manager.Ĭhromatogram files can be easily converted to ABI, FASTA, SEQ or TXT with tools like DNA Sequence Assembler, DNA The green bars above chromatogram peaks high confidence scores.ĭNA Sequence Assembler supports SCF and ABI (ABI, AB, AB!, AB1) chromatogram files. A chromatogram (sometimes also called electropherogram) is the visual representation of a DNA sample produced by a sequencing machine (such as Applied Biosystems ABI PRISM 7700 Sequenceįig. This post is essentially a plea to the internet for someone to tell me there's a better program/method out there!. This combination is fairly straightforward when the sequencing confirms everything nicely, but when there's been a problem somewhere it takes a lot of time and effort to highlight it. Reverse Complement ( ) - for RCing sequencesĬlustal Omega ( ) - to align my sequencing to the sequences I have in my maps and check it's correct or notĮxPASy ( ) - to check the sequenced data is 'in frame' and that the translated protein sequence is how it should beīLAST ( ) - last resort in bizarre cases of "wtf is this random sequence in the middle of things?" Microsoft Word - to handle raw/FASTA sequences Snapgene (free) viewer - for plasmid map creation/visualisation/restriction site mapping/chromatogram viewer My question is: What programs do people recommend for looking at your sequencing data? It's my first time experiencing cloning without being babysat really and the struggle appears with the sequencing data. Creating a few simple constructs of fluorescent reporter tagged proteins. 
I'm a first year PhD Biochemistry student and I've been doing some cloning recently, fairly simple stuff, just ligating sequences into a 3.1 hygro vector. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |

 RSS Feed
RSS Feed
